Dear SCT users,
We are happy to announce a new SCT release!
This is a beta version for now, until we gather your feedback and fix a few remaining issues.
Changes to release
Installation instruction
If you have any question or feature request, please post in this forum.
Happy processing!
The Spinal Cord Toolbox Team
Hi Julien,
I wanted to let you know that I am experiencing difficulties installing v4.0.0-beta.2 on two of my three Macs. The installations were attempted via the provided command:
git clone --branch=master https://github.com/neuropoly/spinalcordtoolbox.git sct
The installation was successful on my laptop – currently running 10.12.6 – but it did not work on my home and work desktops running 10.14.3 and 10.14.4, respectively. I have attached text files for potential debugging purposes. Also, I was concerned that the installation potentially failed on my work computer due to conflict(s) with other software that has been installed over the years, so this installation of the SCT was attempted after a fresh wipe and reinstall of the OS.
Please let me know if there’s anything that I can do to help troubleshoot. Thank you!
Kind regards,
-Rob
SCT installation failed on macOS 10_14_4 on 2019-04-19.txt (109.4 KB)
SCT installation failed on macOS 10_14_3 on 2019-04-22.txt (110.3 KB)
Thank you for the feedback @barryrl!
Yes, we are aware of this issue (ticket opened here). We will fix it today 
Cheers
Julien
1 Like
Hi Julien,
I also face a problem with the new version. I ran a git pull and now the error message says something about a missing module named portalocker.
I ran the sct_check_dependencies because I thought that it might install this missing python package but unfortunately, this command gives the same error message:
Spinal Cord Toolbox (master/4c2aca38d2536e202d58b9be646652a0f4280b6a)
Running /home/tinnermann/sct/scripts/sct_check_dependencies.py
Traceback (most recent call last):
File “/home/tinnermann/sct/scripts/sct_check_dependencies.py”, line 32, in
import sct_utils as sct
File “/home/tinnermann/sct/scripts/sct_utils.py”, line 26, in
import portalocker
ImportError: No module named portalocker
Total processing time: 0 min 1 s
Should I reinstall the toolbox or manually install portalocker?
Best,
Alexandra
Hi Alexandra,
Indeed, you are missing a Python library that has recently been added. However, with the recent move to Python 3, I suggest you do a fresh install:
- Remove your sct folder
- git clone sct
- run installer
Let me know how it goes,
Julien
Hi Julien,
I missed the info about the move to python3. With the fresh install everythings seems to work fine.
Thanks for your fast reply!
By the way, I now have a mean epi image of 82 participants that looks pretty amazing!!
Thanks for all your help!
Best
Alexandra
Awesome! Thanks for the feedback
Hi Julien,
sorry for bothering you again…
I noticed a difference between my previous version and beta.2/beta.3 concerning the field of view of my mean epi after registering it to template space. In the previous version, the field of view of the mean epi was larger than with the new version although image dimensions are identical.
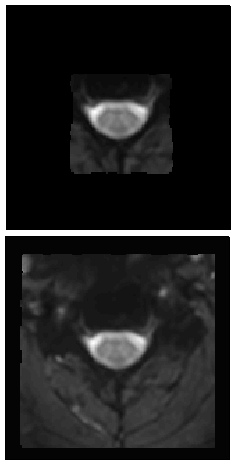
Do you have any explanation for that or any idea why this happens?
I tried to find the point in my analysis that somehow provokes this difference by running different analysis steps with different versions. It seems that if I run the coregistration from my t1 image to template space with the old version and all other analysis steps with the new version, the difference in fov does not occur. That somehow tells me that the warp_t12template (which I also use as initwarp for my epi to template registration) should differ between the versions. Still, I am wondering why the t1 warp should cause the smaller fov. Anything else I could check?
Best,
Alexandra
Dear Alexandra,
You are right, this is a regression bug. A ticket has been opened here and will be addressed with top priority.
Thank you very much for letting us know about this.
Hi Alexandra,
We fixed the issue in 4.0.0-beta.4.
Thank you for the feedback!
Julien
Cool, thanks a lot Julien!

