I would like to know which is the best approach to deal with spinal cord data undergoing DKI analysis.
Here you find attached the DKI dataset. The aim is removing artifacts and computing DKI main metrics.
Thank you in advance,
Rosella Trò
https://drive.google.com/open?id=12eb3eJL_RWmggxT3TQ2AEQ28zkVnXk3z
Hi @Rosella_Tro,
The best approach would be to do all the processing exemplified by the batch_processing.sh example script, and only replace the function sct_dmri_compute_dti with your favourite DKI analysis software.
Best,
Julien
Thank you very much,
Rosella
One more thing, is it possible for me to use batch_processing.sh if I just have data acquired for DKI analysis and not T1 and T2 ones?
hi,
You could re-use part of it. It depends on the scenario (what you would like to measure, what data you have, what prior in terms of slice localization, etc.). For example, if you know that your DKI slices are centered at a particular level, then you can use a single-point registration to the template, as illustrated in the SCT course (see p74 of the Beijing SCT course).
Best,
Julien
Hi,
what I need to do is cleaning data from artifacts and create DKI parametric maps.
Please find attached an example of my data.
https://drive.google.com/open?id=12eb3eJL_RWmggxT3TQ2AEQ28zkVnXk3z
Hi,
If you only need to create DKI maps and extract metrics, you could do something like this (replace ${file_dwi} with your file name):
# Separate b=0 and DW images
sct_dmri_separate_b0_and_dwi -i ${file_dwi}.nii.gz -bvec ${file_bvec}
# Segment cord (1st pass -- just to get a rough centerline)
sct_propseg -i ${file_dwi}_dwi_mean.nii.gz -c dwi
# Create mask to help motion correction and for faster processing
sct_create_mask -i ${file_dwi}_dwi_mean.nii.gz -p centerline,${file_dwi}_dwi_mean_seg.nii.gz -size 30mm
# Crop data for faster processing
sct_crop_image -i ${file_dwi}.nii.gz -m mask_${file_dwi}_dwi_mean.nii.gz -o ${file_dwi}_crop.nii.gz
# Motion correction
sct_dmri_moco -i ${file_dwi}_crop.nii.gz -bvec ${file_dwi}.bvec -x spline
file_dwi=${file_dwi}_crop_moco
file_dwi_mean=${file_dwi}_dwi_mean
# Segment spinal cord (only if it does not exist)
sct_deepseg_sc -i ${file_dwi_mean} -c dwi -qc qc
file_dwi_seg=${file_dwi_mean}_seg
# Create disc label, assuming that the FOV is centered at the C2/C3 disc
# WARNING: You should adapt the code below if your FOV is centered somewhere else
sct_label_utils -i ${file_dwi_seg} -create-seg -1,3 -i labels_disc.nii.gz
# Register template->dwi
sct_register_to_template -i ${file_dwi_mean}.nii.gz -s ${file_dwi_seg}.nii.gz -ldisc labels_disc.nii.gz -ref subject
# Warp template
sct_warp_template -d ${file_dwi_mean}.nii.gz -w warp_template2anat.nii.gz -qc qc
# Create mask around the spinal cord (for faster computing of DTI maps)
sct_maths -i ${file_dwi_seg}.nii.gz -dilate 3,3,3 -o ${file_dwi_seg}_dil.nii.gz
# Compute DTI using RESTORE
sct_dmri_compute_dti -i ${file_dwi}.nii.gz -bvec ${file_bvec} -bval ${file_bval} -method standard -m ${file_dwi_seg}_dil.nii.gz
# ADD YOUR DKI CALCULATION HERE
# Compute FA, MD and RD in WM between C2 and C5 vertebral levels
sct_extract_metric -i dti_FA.nii.gz -f label/atlas -l 51 -vert 2:5 -o ${PATH_RESULTS}/DWI_FA.csv -append 1
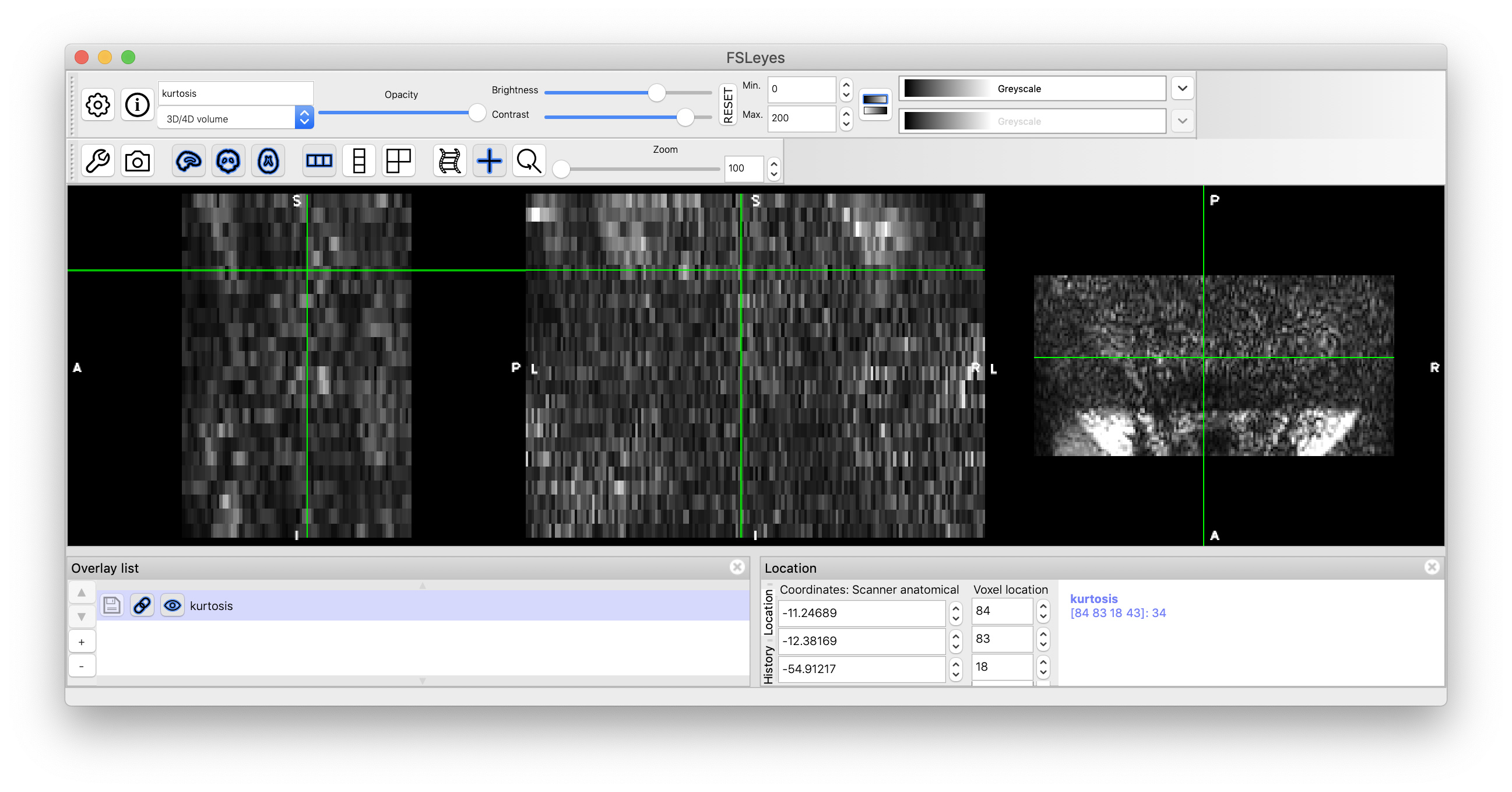
Please note however that your data seem to have some issues. There is almost no visible contrast across DW images in the spinal cord. Related to that, it seems like a saturation band was applied directly on the spinal cord. See the screenshot below (the green cross is centered in the spinal cord):
Best,
Julien
(post withdrawn by author, will be automatically deleted in 24 hours unless flagged)
Hi,
I know my data are not so good but they come from children and we need to keep the acquisition time as short as possible.
I will use the script you provide me to compute DKI metrics.
I also have pediatric data from 1 or 2 month-neonates, I have tried to preprocess them with SCT but even the crop function does not seem to work. They are cervical spinal cord data.
Please find attached a link to these data. If you could help me to decide how to handle them with SCT I would be very grateful.
Thank you,kurtosis_crop_moco.bval (224 Bytes)
kurtosis_crop_moco.bvec (1.8 KB)
https://onedrive.live.com/?id=B0C0561E34E9D76F!586&cid=B0C0561E34E9D76F
Hi @Rosella_Tro,
The one drive link asks me for a username/password.
Also, it would be helpful if you could let me know the syntax you used, when you say “the crop function does not seem to work”. I will be in a better position to instruct as to why it doesn’t work, and suggest proper syntax.
Best,
Julien
Hi,
Ok I will try to be more clear. I am using the commands suggested in your course to pre/process and crop dMRI data in order to subsequently compute both DTI and DKI data. I will attach both the script I use. The commands fail presumably because it is cervical spinal cord from neonates, so the anatomical size of the district is very small and also part of the brain is acquired. As a result, the crop function crops something which does not corresponds to the SC. The same procedure seemed to work better with children, as the spinal cord is bigger than newborns’.
Thank you ,
Rosella
PS: Unfortunately, I am unable to attach my data as they are too heavy, and if I try and attach a Google Drive link, my post get hidden by the forum itself.
script_soggetti_spinale.sh (6.2 KB)
kurtosis_crop_moco.bval (224 Bytes)
kurtosis_crop_moco.bvec (1.8 KB)
Hi
I recovered your post with the gdrive link (it was flagged as spam by the system).
I will download your data and have a look when I find some time, hopefully today
It would be great,
thank you,
Rosella
@Rosella_Tro the drive link is not there anymore. Could you please put it back? I removed the filter that spams posts with drive links in there.
sorry about the inconvenience
sorry for the delay in my answer but I have data in my office lab. I have tried to attach the google drive link but the post gets flagged by the community again. do you manage to recover that link from the spam?
thank you,
Rosella
Sorry it doesn’t seem to work— can you just email me the link?
@Rosella_Tro I did have a look at your data and came up with a dedicated workflow for this dataset, where part of the image should be ignored from the analysis. I’ve created a video tutorial which you can access here.
Below is the series of script that I’ve used for your data. As you can see, the sct_propseg line was replaced with sct_deepseg_sc, and includes manual initialization:
sct_dmri_separate_b0_and_dwi -i kurtosis.nii -bvec kurtosis_crop_moco.bvec
sct_deepseg_sc -i kurtosis_dwi_mean.nii -c dwi -centerline viewer
sct_create_mask -i kurtosis_dwi_mean.nii -p centerline,kurtosis_dwi_mean_seg.nii -size 30mm -o mask_kurtosis_dwi_mean.nii
sct_crop_image -i kurtosis.nii -m mask_kurtosis_dwi_mean.nii -o kurtosis_crop.nii
sct_dmri_moco -i kurtosis_crop.nii -bvec kurtosis_crop_moco.bvec -bval kurtosis_crop_moco.bval -x spline
sct_deepseg_sc -i kurtosis_crop_moco_dwi_mean.nii -c dwi
Hope that helps.
Best,
Julien
I expected I should use a manual initialization and I will try to fix the viewer function on my computer as probably some library is not installed. Anyway, thank you very much for the help. If you wish, I will update you with my results .
Thank you,
Rosella
When runnign the script you provide me, the following error raises:
Spinal Cord Toolbox (dev)
Config deepseg_sc:
_ Centerline algorithm: viewer_
_ Brain in image: False_
_ Kernel dimension: 2d_
_ Contrast: dwi_
_ Threshold: 0.01_
Create temporary folder (/tmp/sct-20191113102740.638922-xrk1ttx)…_
Reorient the image to RPI, if necessary…
Finding the spinal cord centerline…
Traceback (most recent call last):
_ File “/home/rosella/sct_dev/scripts/sct_deepseg_sc.py”, line 201, in _
_ main()_
_ File “/home/rosella/sct_dev/scripts/sct_deepseg_sc.py”, line 185, in main_
_ threshold_seg=threshold, remove_temp_files=remove_temp_files, verbose=verbose)_
_ File “/home/rosella/sct_dev/spinalcordtoolbox/deepseg_sc/core.py”, line 477, in deep_segmentation_spinalcord_
_ centerline_fname=file_ctr)_
_ File “/home/rosella/sct_dev/spinalcordtoolbox/deepseg_sc/core.py”, line 106, in find_centerline_
_ im_labels = call_viewer_centerline(im)
_ File “/home/rosella/sct_dev/spinalcordtoolbox/centerline/core.py”, line 268, in call_viewer_centerline
_ from spinalcordtoolbox.gui.base import AnatomicalParams_
_ File “/home/rosella/sct_dev/spinalcordtoolbox/gui/base.py”, line 18, in _
_ from PyQt5 import QtCore, QtGui, QtWidgets_
ImportError: dlopen: cannot load any more object with static TLS
–
I am currently searching how to solve it.
Thank you,
Rosella
hum… this might be a memory issue (might be related to this).
could you please run the following commands and copy/paste the output:
sct_check_dependencies
sct_testing