Dear Experts
Is it possible to do the following:
1, Have two large scans (~250x250x18 voxels in-plane); A and B
2, Segment them to get the centerline for better coregistration (A_seg and B_seg) and for masking
3, Mask and crop them both with the same mask (cutting it down to 80x80x18 voxels); A_crop and B_crop
4, Coregister the two masked scans (A_crop to B_crop)
5, Apply the warp to the original A?
(This is a follow-up from an old question of mine: Non-rigid coregistration shrinks cord (sct_register_multimodal, syn algo, 4.0 beta 2))
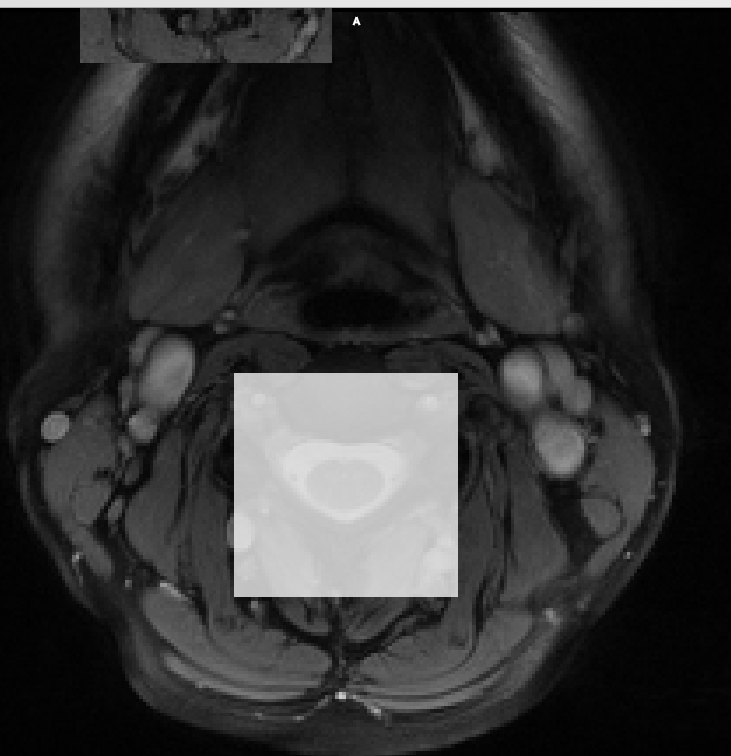
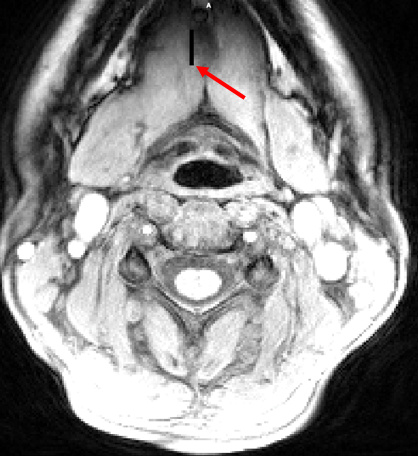
Steps 1-4 work fine for me, but when I apply the warp to the uncropped image, the end result is confusing, and the warp seems to be applied not to the cropped and masked area, but to a corner of the original image. See the black tear in the top of this image
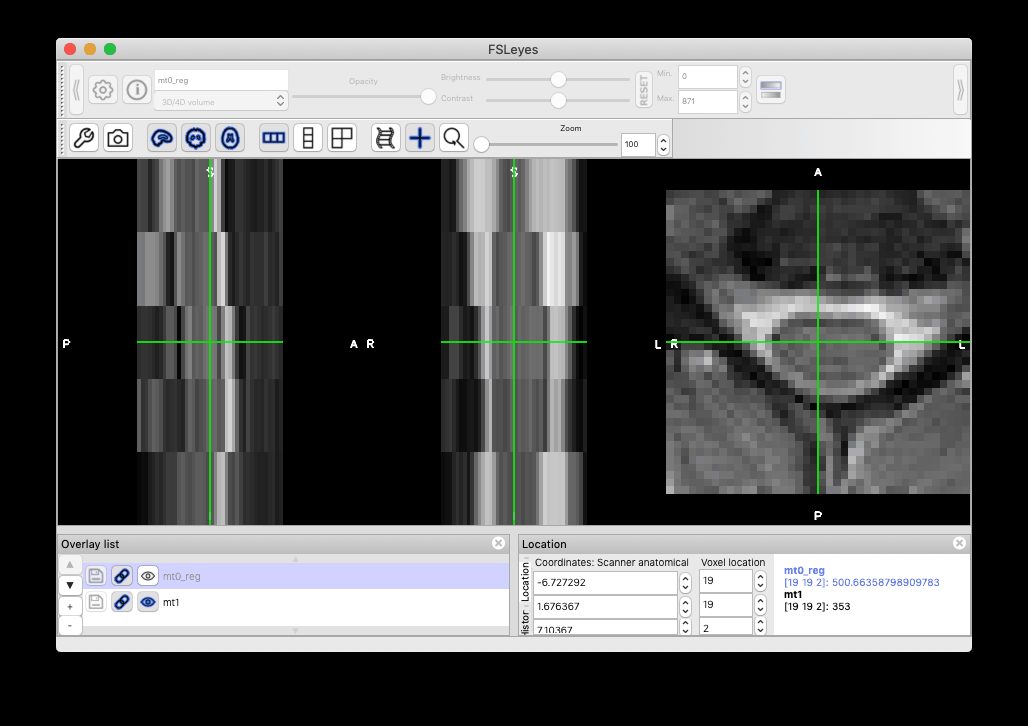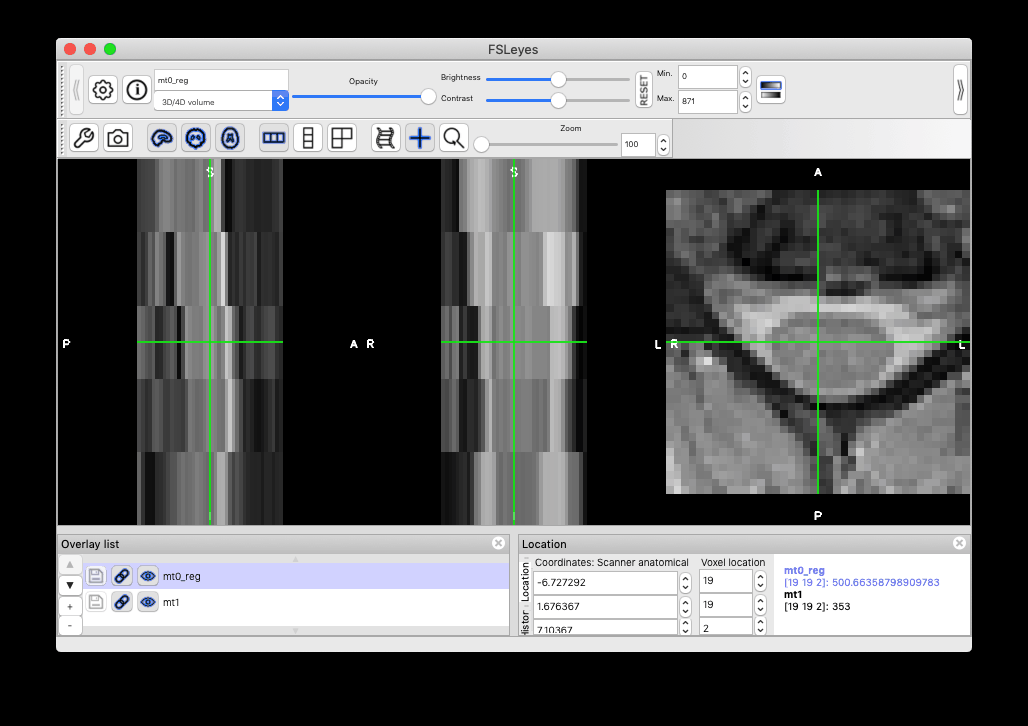
I think it is down to the cooridante system of the cropped scans and the uncropped ones not being the same. When visualised in FSLeyes, the cropped scan is in the upper corned of it’s own uncropped version (original scan in full view, mask in white opaque overlay, cropped scam in the top, close to the left edge):
I am guessing the way around this is to properly define the -d destination files. Here is my code, using these mock filenames.
Coregistration
‘sct_register_multimodal -i A_crop.nii -iseg A_seg.nii -d B_crop.nii -dseg B_seg.nii -param step=1,type=seg,algo=centermassrot:step=2,algo=bsplinesyn,type=im,metric=MI,iter=3,slicewise=0 -o B_reg.nii -owarp A_warptoB.nii -qc /QAfolder’
Warping
sct_apply_transfo -i A.nii -d B.nii -w A_warptoB.nii
SCT version 4.2. I tried checking pages 99-101 of the most recent SCT course (20190122), but it appears I am doing things right.
What am I doing wrong?