Hi,
I would like to know which warping fields (/tmp//warp) should I use to get a straight/LR symmetrical SC in PAM50 space (Step 1 to Step 5) without the non-rigid xy transformation (Step 6) and with only 1 interpolation?
Thank you,
Mounir M El Mendili
Hi Mounir,
If you keep the temporary folders on running sct_register_to_template (using flag -r 0), go to the temp folder, which is indicated in the STDOUT of your Terminal, example (see last line):
julien-macbook:~/sct_course_data/single_subject/data/t2 $ sct_register_to_template -i t2.nii.gz -s t2_seg.nii.gz -ldisc t2_labels_vert.nii.gz -c t2 -r 0
--
Spinal Cord Toolbox (git-master-3ce19094c99ea201fdc6757792e566ca3d7a9bb8)
sct_register_to_template -i t2.nii.gz -s t2_seg.nii.gz -ldisc t2_labels_vert.nii.gz -c t2 -r 0
--
Check template files...
OK: /Users/julien/code/spinalcordtoolbox/data/PAM50/template/PAM50_t2.nii.gz
OK: /Users/julien/code/spinalcordtoolbox/data/PAM50/template/PAM50_label_disc.nii.gz
OK: /Users/julien/code/spinalcordtoolbox/data/PAM50/template/PAM50_cord.nii.gz
Check parameters:
Data: t2.nii.gz
Landmarks: t2_labels_vert.nii.gz
Segmentation: t2_seg.nii.gz
Path template: /Users/julien/code/spinalcordtoolbox/data/PAM50
Remove temp files: 0
Check input labels...
Creating temporary folder (/var/folders/qd/2g7h7921023d_rzm472bf8jm0000gn/T/sct_2025-12-10_11-54-25_register-to-template_mt5dw3w0)
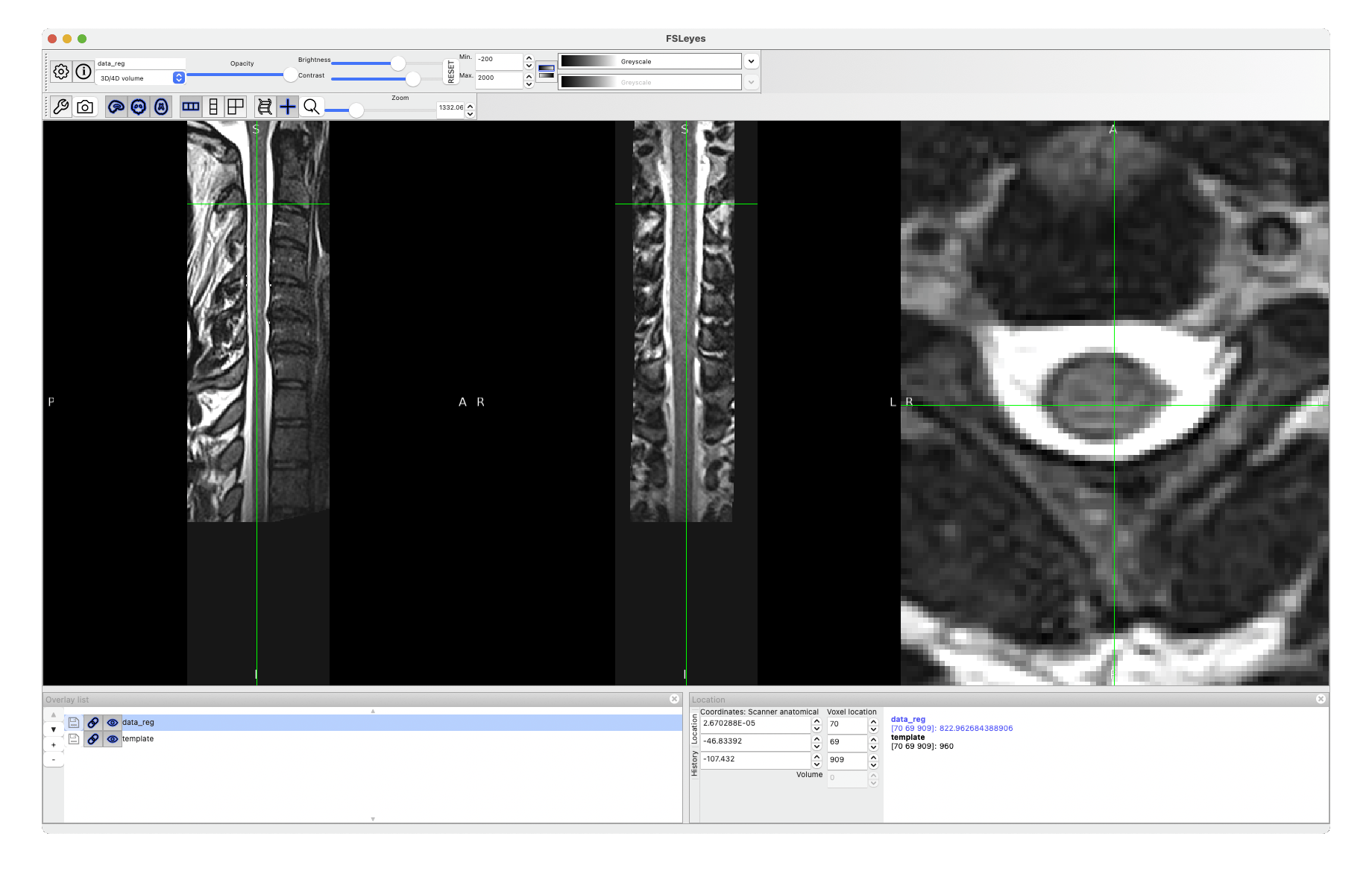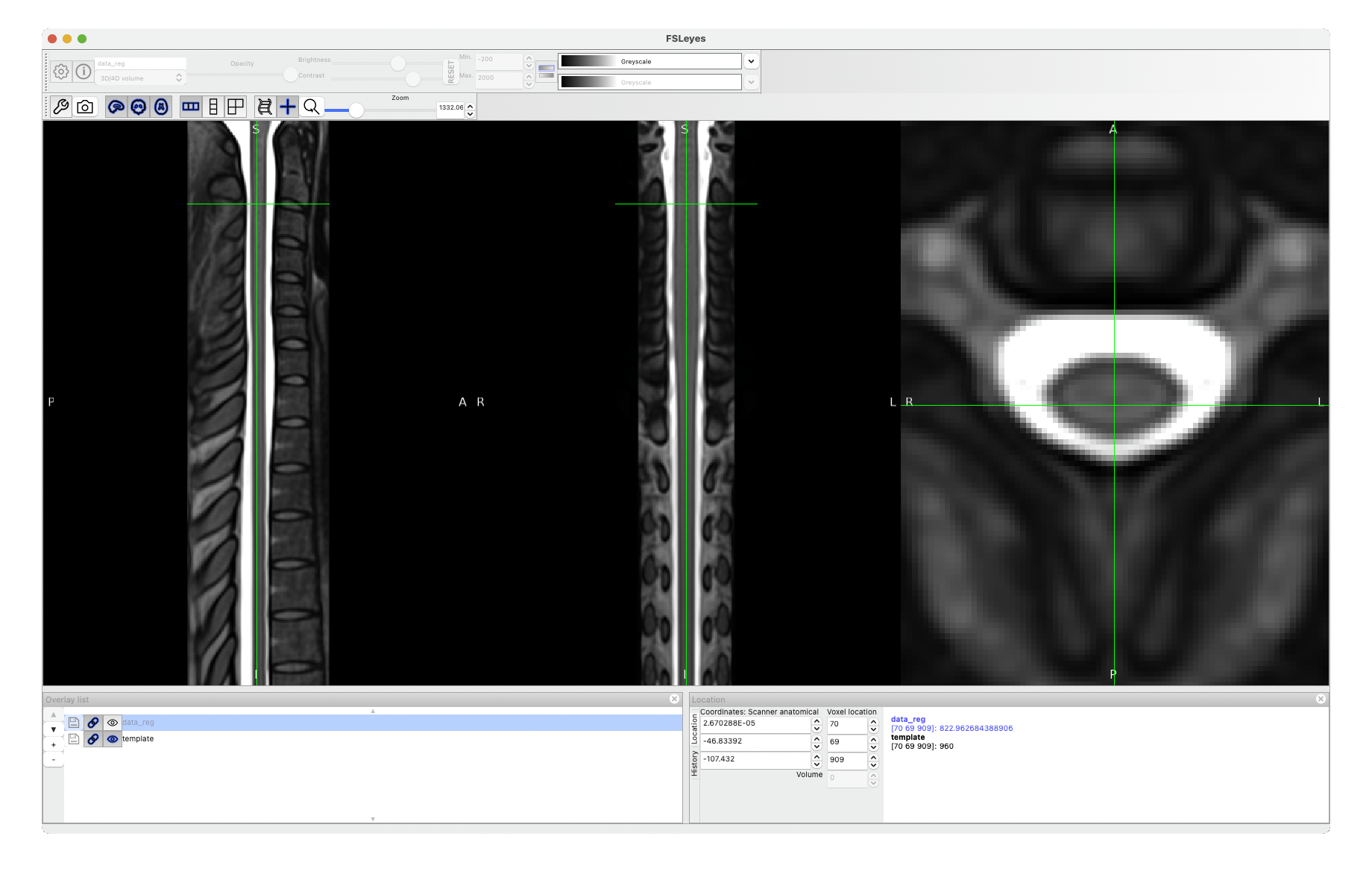
The warping field you’re interested in is called warp_curve2straightAffine.nii.gz.
See what it produces after running:
sct_apply_transfo -i data.nii -d template.nii -w warp_curve2straightAffine.nii.gz
Cheers,
Julien
I’m trying to figure out which warp field should I use to transform the PAM50 template into the curve2straightAffine space. My current assumption is to use: warp_template_crop_sub2data_rpi_1mm_straightAffine_crop_sub.nii.gz
when I apply this warp, the overlap between warped PAM50 and curve2straightAffine_t2w is not that good. The terminal instructions seem straightforward, but I’m not sure if I’m interpreting them correctly?
Thank you very much!
Mounir
warp_template_crop_sub2data_rpi_1mm_straightAffine_crop_sub.nii.gz is a temporary warping field that uses a cropped version of the PAM50 template, which is done to accelerate processing. So no, you cannot apply that warping field on the PAM50 native template.
Maybe you can tell us exactly what you are trying to achieve?
I would prefer to analyze my cohort in an intermediate/common space rather than using the PAM50 space, because the following SyN registration would apply a very strong deformation on my images.
→ I want to bring the PAM50 template to that common space so I can use WM/GM probabilistic segmentations and WM atlases.
Is it possible to have the cropped version of the PAM50 template/the cropping window coordinates?
Thanks for your help!
Mounir
If you just want to skip the last non-linear deformations, you can simply customize the flag -param to specify only one step without iteration (equivalent of just running step=0), example:
sct_register_to_template -i t2.nii.gz -s t2_seg.nii.gz -l t2_labels_vert.nii.gz -c t2 -param step=1,type=seg,algo=syn,iter=0
Hi Julien,
I tried what you proposed, and it worked. Thank you very much!
I also tested rootlets-informed registration to the PAM50 template. My T2w was segmented accurately, and the rootlets were also segmented and labeled accurately.
However, when I tried to bring my segmentation into PAM50 space, a small portion of the segmentation is normalized inaccurately. The most surprising part is that the normalized T2w image looks fine (see attached).
This issue occurred in only few subjects in my dataset.
Here are the command lines I used:
sct_register_to_template -i t2.nii.gz -s t2_seg.nii.gz -lrootlet t2_rootlets.nii.gz -ofolder ./ -c t2 -param step=1,type=segm,algo=syn,iter=0 # I also tried “type=rootlet”
sct_apply_transfo -i t2_seg.nii.gz -d PAM50_t2.nii.gz -w warp_curve2straight.nii.gz warp_anat2template.nii.gz -o t2_seg_PAM50.nii.gz -x nn
sct_apply_transfo -i t2_rootlets.nii.gz -d PAM50_t2.nii.gz -w warp_curve2straight.nii.gz warp_anat2template.nii.gz -o t2_rootlets_PAM50.nii.gz -x nn
Would you have any suggestions on what might be causing this issue?
Best,
Mounir
Hi Mounir,
Could you please share the files t2_seg.nii.gz and t2_rootlets.nii.gz for one of the subjects where you see this issue?
Julien
Thanks for your answer. Please find attached the files t2_seg.nii.gz and t2_rootlets.nii.gz.
t2_rootlets.nii.gz (34.0 KB)
t2_seg.nii.gz (11.4 KB)
Hi Mounir,
I tried running the registration using the t2_seg as the input image (since I don’t have access to it), and the registration looks OK to me:
This is what I ran:
sct_register_to_template -i t2_seg.nii.gz -s t2_seg.nii.gz -lrootlet t2_rootlets.nii.gz -ofolder ./ -c t2 -param step=1,type=seg,algo=syn,iter=0
So I suspect something is off with your T2w image. You can email it to me if you wish.
P.S. Please note that there is a typo in your syntax:
type=segm should be type=seg.
Actually, when applying the output warping field to the PAM50_seg image, I can reproduce the issue:
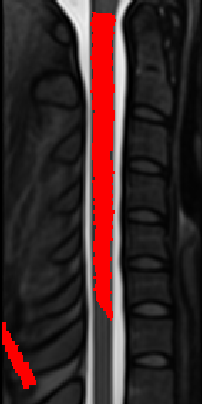
Full code:
sct_register_to_template -i t2_seg.nii.gz -s t2_seg.nii.gz -lrootlet t2_rootlets.nii.gz -ofolder ./ -c t2 -param step=1,type=seg,algo=syn,iter=0 -qc qc
sct_apply_transfo -i t2_seg.nii.gz -d $SCT_DIR/data/PAM50/template/PAM50_t2.nii.gz -w warp_curve2straight.nii.gz warp_anat2template.nii.gz -o t2_seg_PAM50.nii.gz -x nn
fsleyes $SCT_DIR/data/PAM50/template/PAM50_t2.nii.gz t2_seg_PAM50.nii.gz &
One possible avenue, is the warning I received when running sct_register_to_template:
Image /private/var/folders/qd/2g7h7921023d_rzm472bf8jm0000gn/T/sct_2026-04-21_13-58-03_register-to-template_5dxpqy_d/seg_bin_rpi_1mm_crop.nii.gz has different qform and sform matrices. This can produce incorrect results. Please use ‘sct_image -i /private/var/folders/qd/2g7h7921023d_rzm472bf8jm0000gn/T/sct_2026-04-21_13-58-03_register-to-template_5dxpqy_d/seg_bin_rpi_1mm_crop.nii.gz -header’ to check that both affine matrices are valid. Then, consider running either ‘sct_image -set-sform-to-qform’ or ‘sct_image -set-qform-to-sform’ to fix any discrepancies you may find.
Possibly related to the issue. I’m investigating… ![]()
OK, I got it. Your syntax in this post was incorrect:
sct_apply_transfo -i t2_seg.nii.gz -d PAM50_t2.nii.gz -w warp_curve2straight.nii.gz warp_anat2template.nii.gz -o t2_seg_PAM50.nii.gz -x nn
You’re concatenating warp_curve2straight and warp_anat2template, but the latter transformation already includes warp_anat2template, so you’re essentially applying it twice. The correct syntax is:
sct_apply_transfo -i t2_seg.nii.gz -d $SCT_DIR/data/PAM50/template/PAM50_t2.nii.gz -w warp_anat2template.nii.gz -o t2_seg_PAM50.nii.gz -x nn
Which should then output:

Please follow the official example batch script provided in SCT’s website. Also go through the SCT course for more details on these processes.
An additional consideration: You’re using -lrootlet, which, for step=0, aligns all rootlets between the native space and the PAM50 space. Since rootlets only go down to C8 spinal level, everything below it is cropped out for subsequent registration steps, hence, the final “native ↔ PAM50” warping field will ignore everything below the C8 spinal level. This is indicated in the sct_register_to_template help:
If more than two labels are used, a non-linear registration will be applied to align the each intervertebral disc between the subject and the template, as described in sct_straighten_spinalcord. This the most accurate method, however it has some serious caveats:
- This feature is not compatible with the parameter
-ref subject, where only a rigid registration is performed.- Due to the non-linear registration in the S-I direction, the warping field will be cropped above the top label and below the bottom label. Applying this warping field will result in a strange-looking registered image that has the same value above the top label and below the bottom label. But if you are not interested in these regions, you do not need to worry about it.
If you’re interested in levels below, consider using disc segmentation instead.
I see. Since sct_apply_transfo relies on ANTs registration, I thought I needed to concatenate the two transformations.
Thank you very much for your help!